Research led by a team of scientists from the Texas A&M School of Veterinary Medicine and Biomedical Sciences puts to bed the heated scientific debate regarding the history of mammal diversification as it relates to the extinction of the non-avian dinosaurs. Their work provides a definitive answer to the evolutionary timeline of mammals throughout the last 100 million years.
The study, published in Science, is part of a series of articles released by the Zoonomia Project, a consortium of scientists from around the globe that is using the largest mammalian genomic dataset in history to determine the evolutionary history of the human genome in the context of mammalian evolutionary history. Their ultimate goal is to better identify the genetic basis for traits and diseases in people and other species.
The Texas A&M University research—led by Dr. William J. Murphy, a professor in the Department of Veterinary Integrative Biosciences, and Dr. Nicole Foley, an associate research scientist in Murphy’s lab—is rooted in phylogeny, a branch of biology that deals with the evolutionary relationships and diversification of living and extinct organisms.
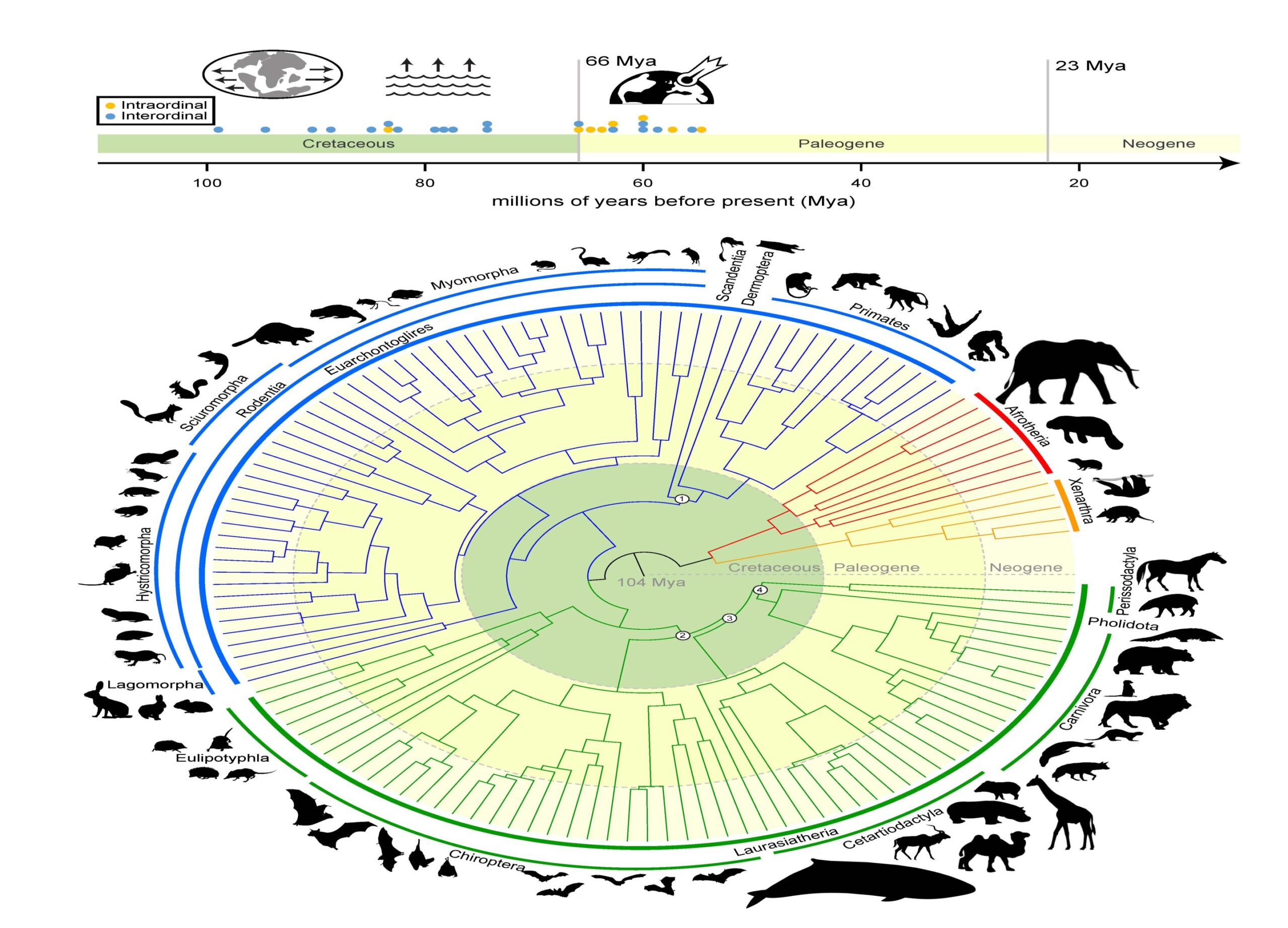
“The central argument is about whether placental mammals (mammals that develop within placentas) diverged before or after the Cretaceous-Paleogene (or K-Pg) extinction event that wiped out the non-avian dinosaurs,” Foley shared. “By performing new types of analyses only possible because of Zoonomia’s massive scope, we answer the question of where and when mammals diversified and evolved in relation to the K-Pg mass extinction.”
The research—which was conducted with collaborators at the University of California, Davis; University of California, Riverside; and the American Museum of Natural History—concludes that mammals began diversifying before the K-Pg extinction as the result of continental drifting, which caused the Earth’s land masses to drift apart and come back together over millions of years. Another pulse of diversification occurred immediately following the K-Pg extinction of the dinosaurs, when mammals had more room, resources and stability.
This accelerated rate of diversification led to the rich diversity of mammal lineages—such as carnivores, primates and hoofed animals—that share the Earth today.
Murphy and Foley’s research was funded by the National Science Foundation and is one part of the Zoonomia Project led by Elinor Karlsson and Kerstin Lindblad-Toh, of the Broad Institute, which also compares mammal genomes to understand the basis of remarkable phenotypes—the expression of certain genes such as brown vs. blue eyes—and the origins of disease.
Foley pointed out that the diversity among placental mammals is exhibited both in their physical traits and in their extraordinary abilities.
“Mammals today represent enormous evolutionary diversity—from the whizzing flight of the tiny bumblebee bat to the languid glide of the enormous Blue Whale as it swims through Earth’s vast oceans. Multiple species have evolved to echolocate, some produce venom, while others have evolved cancer resistance and viral tolerance,” she said.
“Being able to look at shared differences and similarities across the mammalian species at a genetic level can help us figure out the parts of the genome that are critical to regulate the expression of genes,” she continued. “Tweaking this genomic machinery in different species has led to the diversity of traits that we see across today’s living mammals.”
Murphy shared that Foley’s resolved phylogeny of mammals is crucial to the goals of the Zoonomia Project, which aims to harness the power of comparative genomics as a tool for human medicine and biodiversity conservation.
“The Zoonomia Project is really impactful because it’s the first analysis to align 241 diverse mammalian genomes at one time and use that information to better understand the human genome,” he explained. “The major impetus for putting together this big data set was to be able to compare all of these genomes to the human genome and then determine which parts of the human genome have changed over the course of mammalian evolutionary history.”
Determining which parts of genes can be manipulated and which parts cannot be changed without causing harm to the gene’s function is important for human medicine. A recent study in Science Translational Medicine led by one of Murphy and Foley’s colleagues, Texas A&M geneticist Dr. Scott Dindot, used the comparative genomics approach to develop a molecular therapy for Angelman syndrome, a devastating, rare neurogenetic disorder that is triggered by the loss of function of the maternal UBE3A gene in the brain.
Dindot’s team took advantage of the same measures of evolutionary constraint identified by the Zoonomia Project and applied them to identify a crucial but previously unknown genetic target that can be used to rescue the expression of UBE3A in human neurons.
Murphy said expanding the ability to compare mammalian genomes by using the largest dataset in history will help develop more cures and treatments for other species’ ailments rooted in genetics, including cats and dogs.
“For example, cats have physiological adaptations rooted in unique mutations that allow them to consume an exclusively high-fat, high-protein diet that is extremely unhealthy for humans,” Murphy explained. “One of the beautiful aspects of Zoonomia’s 241-species alignment is that we can pick any species (not just human) as the reference and determine which parts of that species’ genome are free to change and which ones cannot tolerate change. In the case of cats, for example, we may be able to help identify genetic adaptations in those species that could lead to therapeutic targets for cardiovascular disease in people.”
Murphy and Foley’s phylogeny also played an instrumental role in many of the subsequent papers that are part of the project.
“It’s trickle-down genomics,” Foley explained. “One of the most gratifying things for me in working as part of the wider project was seeing how many different research projects were enhanced by including our phylogeny in their analyses. This includes studies on conservation genomics of endangered species to those that looked at the evolution of different complex human traits.”
Foley said it was both meaningful and rewarding to definitively answer the heavily debated question about the timing of mammal origins and to produce an expanded phylogeny that lays the foundation for the next several generations of researchers.
“Going forward, this massive genome alignment and its historical record of mammalian genome evolution will be the basis of everything that everyone’s going to do when they’re asking comparative questions in mammals,” she said. “That is pretty cool.”
More information:
Nicole M. Foley et al, A genomic timescale for placental mammal evolution, Science (2023). DOI: 10.1126/science.abl8189
Provided by
Texas A&M University
Citation:
Researchers use genomes of 241 species to redefine mammalian tree of life (2023, April 28)
retrieved 28 April 2023
from https://phys.org/news/2023-04-genomes-species-redefine-mammalian-tree.html
This document is subject to copyright. Apart from any fair dealing for the purpose of private study or research, no
part may be reproduced without the written permission. The content is provided for information purposes only.

